|

|
|
|
Primo Optimum 3.4 Help: Optimum For Gene Expression
About Primo Optimum 3.4: Online Primo Optimum is designed for optimal expression of genes of interest in a host organism (e.g., E. Coli, yeast, plants, or mammalian cell lines). Different species have different codon usage frequencies. For high level expression in a host, optimization of the gene sequence to match the host's codon bias increases yield and is frequently necessary. Primo Optimum provides a tool for optimizing gene expression in any host species and designs primers for the one-step or stepwise PCR synthesis of the optimal gene. In addition it checks for a number of factors that might be detrimental for gene expression. For more PCR primer design tools (standard PCR, multiplex PCR, profile PCR, MSP PCR, degenerate PCR, etc.), please visit Primo Home. Browser requirements:Primo Online runs on the following Java-enabled browser:If you use one of the above browser and you can't run Primo Online, please make sure Java is enabled in your browser. For Internet Explorer, go to Tools/Internet Options, click on Security Settings, scroll down to find Microsoft VM, deselect "Disable Java". Mac OSX 10 IE5.1 has a bug, it does not allow copy/paste into a Java text field, thus you will not be able to import a new sequence. Attempting copy/paste may kill your browser. Stand-alone versions don't require IE to run, thus don't have this problem. |
How to use:
To start using Optimum, copy and paste your sequences into the input window. On Windows you may need to use Ctrl-C (copy) and Ctrl-V (paste). On Macs you may need to use Apple-C (copy) and Apple-V (paste). Numbers and white spaces will be ignored.Both nucleotide and protein sequences can be used as input. Users don't need to specify whether the sequence is nucleotides or amino acids, the program will determine the sequence type automatically. Note degenerate codes for nucleotides are not acceptable, since the protein sequence can not be determined uniquely from degenerate nucleotide sequences. (The program might mistake a nucleotide sequence as a protein sequences if it contains degenerate codes.)
For gene synthesis without optimizing codon usage, check "Disable optimization" or input your DNA sequences in the "optimized sequence" window (and leave the top-left windwon blank).
Engineer restriction sites:
Optimum can be used to add/remove restriction sites from the optimized sequences without altering the protein sequences encoded (silent mutation).
 |
Figure 1.Engineering restriction sites (silent mutation)
Step 1.Select a region where you would like to add a restriction site. You may chose the region where two primers overlap. Or input the specific positions (e.g. 50-100 for nucleotide positions 50 to 100). The position input will be ignored if any one of the primer choices is not "N/A." In another word, to input specific positions, users need to select "N/A" for the two overlapping primers. Step 2.Check "Add restriction site" and select "any" from the restriction enzyme list. Press on "Check" to find a list of restriction sites that can be added without altering the protein sequences encoded. Step 3.Uncheck "any" and select one restriction site. Press "Check" and then "Confirm." You may also need to remove any restriction sites outside the region you selected. To do so, simply check "Remove restriction site" and select the enzyme name from the enzyme list before pressing on "Check." |
Behind the scene:
1. DiagnosisTo run the diagnosis function, you must input a nucleotide sequence in the top-left window. The program will find the longest open reading frame, and check for a number of features such as Kozak translation initiation sequence, Shine Dalgarno sequence, rare codons, etc. It is not intended as an extensive diagnosis, rather serves as a check list to remind researchers of optimization options.
2. Optimum reverse translation
Optimum program will reverse translate a protein sequence to a nucleotide sequence using the "optimal codons," i.e., codons with the highest usage frequencies.
If a nucleotide sequence is the input, the longest open reading frame will be determined, and the ORF will be reverse translated. 5' and 3' untranslated regions will not be optimized.
3. Gene synthesis
To design PCR primers for gene synthesis you may input a protein or DNA sequence in the top-left window and let the program determine the optimal sequence. Alternatively leave the top-left window blank, and input or edit your optimal sequence in the window with a label "Optimal DNA Sequence:".
The program will find a series of overlapping PCR primers for the one-step or stepwise PCR synthesis of genes. The program will not check for primer-primer dimers since the templates (primers also are templates) are in much higher concentration.
There are three options for gene synthesis:
Shown in Figure 2 are several experimental designs for gene synthesis. Note the orientations of the primers are different for different experimental designs.
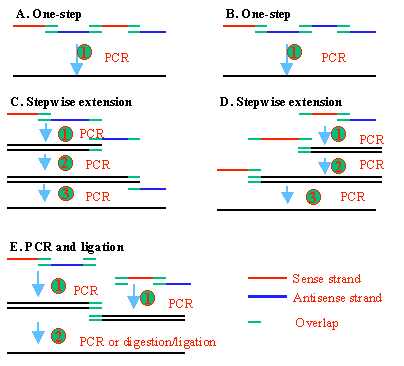
Figure 2. PCR gene synthesis uses a series of overlapping PCR primers, which also serve as templates. In the one-step approach, the outer primers are at a higher concentration than the inner primers. As a result, the full-length gene is eventually synthesized. Primers designed this way may also be used to synthesize the gene fragments using two or three primers at a time and then elongate the PCR product by additional round of PCR synthesis or ligation. Note the orientations of the primers are different for different experimental designs. |
4. Host species
Two species (human and E. coli) are available in the Optimum Online. More species are available in the stand-alone version (see below). Users can add more host species in the stand-alone version, provide that the codon usage frequencies and genetic codes are known for the hosts.
5. Codon usage
The codon usage frequency relative to that of the optimal codon is diagramed. Mouse click on the diagram to show sequences begin at the clicked position. Note the diagram shows the relative codon usage frequencies, while the absolute frequencies are used for the determination of rare codons in the diagnosis mode. The two views complement each other.
6. Add new host species
To add new species, users need to download codon usage table from the Codon Usage Database. Select the species, then the genetic code and the GCG style (Figure 2). The codon usage and genetic code will be combined into one table as shown in Figure 3. Copy the table (without the first row of column names) and use it as the input for the "Add Host" function of Primo Optimum. Alternatively, users may edit the "hostdata.txt" file in the "codontable" folder.

| Figure 2. Codon usage and the genetic codes can be downloaded from the Codon Usage Database. Primo Optimum uses the GCG format |
Gly GGG 7215.00 10.37 0.16 Gly GGA 15218.00 21.86 0.34 Gly GGT 9675.00 13.90 0.21 Gly GGC 12935.00 18.58 0.29 Glu GAG 27761.00 39.88 0.65 Glu GAA 15275.00 21.95 0.35 Asp GAT 15788.00 22.68 0.44 Asp GAC 19762.00 28.39 0.56 Val GTG 18841.00 27.07 0.44 Val GTA 4443.00 6.38 0.10 Val GTT 8958.00 12.87 0.21 Val GTC 10109.00 14.52 0.24 Ala GCG 6471.00 9.30 0.14 Ala GCA 10998.00 15.80 0.24 Ala GCT 13625.00 19.57 0.30 Ala GCC 14418.00 20.71 0.32 Arg AGG 6893.00 9.90 0.18 Arg AGA 9336.00 13.41 0.25 Ser AGT 9043.00 12.99 0.15 Ser AGC 14337.00 20.60 0.24 Lys AAG 20053.00 28.81 0.52 Lys AAA 18383.00 26.41 0.48 Asn AAT 11144.00 16.01 0.38 Asn AAC 18365.00 26.38 0.62 Met ATG 18183.00 26.12 1.00 Ile ATA 4800.00 6.90 0.15 Ile ATT 11105.00 15.95 0.34 Ile ATC 17031.00 24.47 0.52 Thr ACG 5748.00 8.26 0.15 Thr ACA 11238.00 16.15 0.29 Thr ACT 9742.00 14.00 0.25 Thr ACC 12646.00 18.17 0.32 Trp TGG 8046.00 11.56 1.00 End TGA 756.00 1.09 0.49 Cys TGT 7956.00 11.43 0.47 Cys TGC 9101.00 13.08 0.53 End TAG 249.00 0.36 0.16 End TAA 542.00 0.78 0.35 Tyr TAT 8627.00 12.39 0.40 Tyr TAC 12961.00 18.62 0.60 Leu TTG 8148.00 11.71 0.13 Leu TTA 4231.00 6.08 0.07 Phe TTT 11678.00 16.78 0.44 Phe TTC 14848.00 21.33 0.56 Ser TCG 4671.00 6.71 0.08 Ser TCA 9158.00 13.16 0.15 Ser TCT 11737.00 16.86 0.19 Ser TCC 12019.00 17.27 0.20 Arg CGG 4970.00 7.14 0.13 Arg CGA 4720.00 6.78 0.13 Arg CGT 4659.00 6.69 0.12 Arg CGC 7152.00 10.28 0.19 Gln CAG 23603.00 33.91 0.73 Gln CAA 8524.00 12.25 0.27 His CAT 7392.00 10.62 0.39 His CAC 11553.00 16.60 0.61 Leu CTG 24932.00 35.82 0.41 Leu CTA 4105.00 5.90 0.07 Leu CTT 8256.00 11.86 0.13 Leu CTC 11817.00 16.98 0.19 Pro CCG 6863.00 9.86 0.17 Pro CCA 11022.00 15.84 0.27 Pro CCT 11781.00 16.93 0.29 Pro CCC 10427.00 14.98 0.26 | Figure 3. Copy the data in the format shown on the left and use as input for the "Add Host" function of the Primo Optimum. |
Melting temperature is determined by the regions of over-lapping primers for Optimum. Melting temperature is the temperature at which 50% of the oligo and its perfect complement are in duplex. PCR annealing temperature a few degree (4-6) lower than the melting temperature is usually used to increase the probability of primer binding. There are two options for calculating the melting temperature. The first uses the simple rule of 2 degree for each A or T and 4 degree for each C or G.
Melting temperature = 4 * Number of G or C + 2 * Number of A or T.
The second "Nearest N" predicts melting temperature using the "Nearest Neighbor" model (John SantaLucia, Proc. Natl. Acad. Sci. Vol. 95, p1460-1465 (1998)). The cation concentration is assumed to be 50 mM and the primer concentration is assumed to be 200 nanomolar. The "Nearest N" is presented because it is more accurate and other formulae can be viewed as approximations of the "Nearest N".
Species available in the stand-alone version
| Species Name | Common Name |
| Arabidopsis thaliana | Arabidopsis |
| Zea mays | Corn |
| Bos taurus | Cow |
| Drosophila melanogaster | Drosophila |
| E. coli | E. coli |
| Xenopus laevis | Frog |
| Homo sapiens | Human |
| Mus musculus | Mouse |
| Rattus norvegicus | Rat |
| Oryza sativa | Rice |
| Saccharomyces cerevisiae | Yeast |
| Danio rerio | Zebrafish |